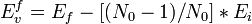Integrated Computational Materials Engineering (ICME)
LAMMPS Vacancy Formation Energy
Abstract
This example shows how to calculate vacancy formation energy (VFE) for FCC metals. The parallel molecular dynamics (MD) code LAMMPS is used to perform the calculations. OVITO is used to visualize the atomic structure before and after the creation of the vacancy. LAMMPS[1].
Author(s): Mohammad Rezaul Karim*, Shakila Taylor*, Firas Akasheh*, Mark A. Tschopp
Advisor(s): Firas Akasheh*, Mark A. Tschopp
(*) Mechanical Engineering Department, Tuskegee University, Tuskegee, AL 36088
Introduction
Typically, metals are crystalline solids that have a periodic crystal structure such as FCC (face centered cubic) and BCC (body centered cubic), both of which are based on the cubic unit cell. The positions of atoms occur in repeating fixed positions, determined by the unit cell parameters. However, the arrangement of atoms in most crystalline materials is not perfect. The regular patterns are interrupted by crystallographic defects. Three types of defects that occur in metals are point, line, and planar defects. In this tutorial we focus on points defects, specifically on vacancies. A vacancy occurs at or around a single lattice point and involves a missing atom. Vacancies are important because they affect the chemical properties of a metal, such as its conductivity, corrosivity, as well as the overall mechanical behavior particularly at high temperatures. An important property of vacancy is its formation energy, Ev. It is the energy required to break the bonds between an atom inside the crystal and its nearest neighbor atoms and removing that atom to where it has no interaction with the remaining system. Another approach puts the removed atom back on the surface of the crystal where some energy is retrieved because new bonds are established with other atoms on the surface. However, there is a net input of energy because there are fewer bonds between surface atoms than between atoms in the interior of the crystal.
Description of Simulation
The LAMMPS first generates a FCC copper (Cu) cell with a 8 x 8 x 8 unit cell dimension and an xyz orientation ([100], [010], and [001], respectively) with a total of 2048 atoms created. The boundaries of the cell structure are periodic in every direction. The input script file removes one atom from the simulation cell and then calculates the vacancy formation energy, Evf, in units of eV after relaxation. The following input script performs the following steps. First, the number of atoms in the cell is counted and the initial energy (Ei) of the system is computed. Then, an atom is removed and the structure is relaxed using energy minimization (conjugate gradient method) and the final energy (Ef) of the relaxed system containing a vacancy is computed. All of the necessary values are stored and are ultimately used to calculate the vacancy formation energy. The equation used to calculate the energy Evf is:
where Ef, Ei, and N0 represent the final energy after the atom is removed from the cell, the initial energy before the atom was removed, and N0 is the total number of atoms in the cell before the vacancy. Once the input file is ran using LAMMPS, the file displays the initial and final energies, the number of atoms, and the vacancy formation energy.
LAMMPS input script for FCC
To run this script, store it in an input file, say "Cu-vacancy.txt" and then use the command "lmp_win_no-mpi.exe < Cu-vacancy.txt" in a Windows environment where "lmp_win_no-mpi.exe" refers to the LAMMPS executable.
# Input file for Vacancy Formation Energy
# --------------- INITIALIZATION ------------------
clear
units metal
dimension 3
boundary p p p
atom_style atomic
# ------------------ ATOM DEFINITION -------------------
variable ao equal 3.615
lattice fcc 3.615
region simbox block -4 4 -4 4 -4 4
create_box 1 simbox
lattice fcc 3.615 orient x 1 0 0 orient y 0 1 0 orient z 0 0 1
create_atoms 1 region simbox
# ------------------------ FORCE FIELDS -----------------------
pair_style eam/alloy
pair_coeff * * FeCuNi.eam.alloy Cu
#---------------------------Settings----------------------------
compute csym all centro/atom fcc
compute eng all pe/atom
compute eatoms all reduce sum c_eng
#----------------------Run Minimization-------------------------
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
dump 1 all custom 400 dump.relax.1.* id type xs ys zs c_csym c_eng
min_style cg
minimize 1e-15 1e-15 5000 5000
run 0
undump 1
#variable N equal count(all), counts the total number of atoms in the cell
#the total number of atoms is stored to the variable N
variable N equal count(all)
variable No equal $N
#variable Ei equal "c_eatoms" computes the initial energy of the cell system before the vacancy
#E is needed to store the initial energy of the system to the variable Ei
variable E equal "c_eatoms"
variable Ei equal $E
#---------------------------------------------------------------
variable r2 equal sqrt(${ao}^2+${ao}^2)/4
#r2 is the radius of the copper atom
#region select is a region defined so that all atoms within this region are removed
region select sphere 0 0 0 ${r2} units box
delete_atoms region select compress yes
#---------------------------------------------------------------------
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
dump 1 all custom 400 dump.relax.2.* id type xs ys zs c_csym c_eng
min_style cg
minimize 1e-15 1e-15 5000 5000
#variable Ef equal "c_eatoms" computes the final energy of the cell system after the vacancy
#The final energy is stored to the variable Ef
variable Ef equal "c_eatoms"
variable Ev equal (${Ef}-((${No}-1)/${No})*${Ei})
#---------------------------------------------
######################################
# SIMULATION DONE
print "All done"
print "Total number of atoms = ${No}"
print "Initial energy of atoms = ${Ei}"
print "Final energy of atoms = ${Ef}"
print "Vacancy formation energy = ${Ev}"
Output
LAMMPS logfile
The log.lammps file should look like the one shown below.
LAMMPS (21 Dec 2011)
# --------------- INITIALIZATION ------------------
clear
units metal
dimension 3
boundary p p p
atom_style atomic
# ------------------ ATOM DEFINITION -------------------
variable ao equal 3.615
lattice fcc 3.615
Lattice spacing in x,y,z = 3.615 3.615 3.615
region simbox block -4 4 -4 4 -4 4
create_box 1 simbox
Created orthogonal box = (-14.46 -14.46 -14.46) to (14.46 14.46 14.46)
1 by 1 by 1 MPI processor grid
lattice fcc 3.615 orient x 1 0 0 orient y 0 1 0 orient z 0 0 1
Lattice spacing in x,y,z = 3.615 3.615 3.615
create_atoms 1 region simbox
Created 2048 atoms
# ------------------------ FORCE FIELDS -----------------------
pair_style eam/alloy
pair_coeff * * FeCuNi.eam.alloy Cu
#---------------------------Settings----------------------------
compute csym all centro/atom fcc
compute eng all pe/atom
compute eatoms all reduce sum c_eng
#----------------------Run Minimization-------------------------
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
dump 1 all custom 400 dump.relax.1.* id type xs ys zs c_csym c_eng
min_style cg
minimize 1e-15 1e-15 5000 5000
WARNING: Resetting reneighboring criteria during minimization (min.cpp:167)
Memory usage per processor = 4.80068 Mbytes
Step PotEng Lx Ly Lz Press Pxx Pyy Pzz eatoms
0 -7250.3671 28.92 28.92 28.92 -86.036792 -86.036792 -86.036792 -86.036792 -7250.3671
1 -7250.3671 28.92 28.92 28.92 -86.036792 -86.036792 -86.036792 -86.036792 -7250.3671
Loop time of 0.0650041 on 1 procs for 1 steps with 2048 atoms
Minimization stats:
Stopping criterion = linesearch alpha is zero
Energy initial, next-to-last, final =
-7250.36709989 -7250.36709989 -7250.36709989
Force two-norm initial, final = 2.85021e-013 2.85021e-013
Force max component initial, final = 7.10543e-015 7.10543e-015
Final line search alpha, max atom move = 0.5 3.55271e-015
Iterations, force evaluations = 1 2
Pair time (%) = 0.028002 (43.0774)
Neigh time (%) = 0 (0)
Comm time (%) = 3.11411e-007 (0.000479064)
Outpt time (%) = 0 (0)
Other time (%) = 0.0370017 (56.9221)
Nlocal: 2048 ave 2048 max 2048 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Nghost: 5765 ave 5765 max 5765 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Neighs: 143360 ave 143360 max 143360 min
Histogram: 1 0 0 0 0 0 0 0 0 0
FullNghs: 286720 ave 286720 max 286720 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Total # of neighbors = 286720
Ave neighs/atom = 140
Neighbor list builds = 0
Dangerous builds = 0
run 0
WARNING: No fixes defined, atoms won't move (verlet.cpp:52)
Memory usage per processor = 4.11404 Mbytes
Step PotEng Lx Ly Lz Press Pxx Pyy Pzz eatoms
1 -7250.3671 28.92 28.92 28.92 -86.036792 -86.036792 -86.036792 -86.036792 -7250.3671
Loop time of 1.13156e-007 on 1 procs for 0 steps with 2048 atoms
Pair time (%) = 0 (0)
Neigh time (%) = 0 (0)
Comm time (%) = 0 (0)
Outpt time (%) = 0 (0)
Other time (%) = 1.13156e-007 (100)
Nlocal: 2048 ave 2048 max 2048 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Nghost: 5765 ave 5765 max 5765 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Neighs: 143360 ave 143360 max 143360 min
Histogram: 1 0 0 0 0 0 0 0 0 0
FullNghs: 286720 ave 286720 max 286720 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Total # of neighbors = 286720
Ave neighs/atom = 140
Neighbor list builds = 0
Dangerous builds = 0
undump 1
#variable N equal count(all), counts the total number of atoms in the cell
#the total number of atoms is stored to the variable N
variable N equal count(all)
variable No equal $N
variable No equal 2048
#variable Ei equal "c_eatoms" computes the initial energy of the cell system before the vacancy
#E is needed to store the initial energy of the system to the variable Ei
variable E equal "c_eatoms"
variable Ei equal $E
variable Ei equal -7250.3671
#---------------------------------------------------------------
variable r2 equal sqrt(${ao}^2+${ao}^2)/4
variable r2 equal sqrt(3.615^2+${ao}^2)/4
variable r2 equal sqrt(3.615^2+3.615^2)/4
#r2 is the radius of the copper atom
#region select is a region defined so that all atoms within this region are removed
region select sphere 0 0 0 ${r2} units box
region select sphere 0 0 0 1.278095507 units box
delete_atoms region select compress yes
Deleted 1 atoms, new total = 2047
#---------------------------------------------------------------------
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
dump 1 all custom 400 dump.relax.2.* id type xs ys zs c_csym c_eng
min_style cg
minimize 1e-15 1e-15 5000 5000
WARNING: Resetting reneighboring criteria during minimization (min.cpp:167)
Memory usage per processor = 4.80068 Mbytes
Step PotEng Lx Ly Lz Press Pxx Pyy Pzz eatoms
0 -7245.5176 28.92 28.92 28.92 -243.32058 -243.32058 -243.32058 -243.32058 -7245.5176
10 -7245.5542 28.92 28.92 28.92 -287.426 -287.426 -287.426 -287.426 -7245.5542
20 -7245.5543 28.92 28.92 28.92 -287.76739 -287.76739 -287.76739 -287.76739 -7245.5543
24 -7245.5543 28.92 28.92 28.92 -287.77273 -287.77273 -287.77273 -287.77273 -7245.5543
Loop time of 0.827048 on 1 procs for 24 steps with 2047 atoms
Minimization stats:
Stopping criterion = linesearch alpha is zero
Energy initial, next-to-last, final =
-7245.51755642 -7245.55428958 -7245.55428958
Force two-norm initial, final = 0.595946 0.000184265
Force max component initial, final = 0.117627 7.40404e-006
Final line search alpha, max atom move = 0.25 1.85101e-006
Iterations, force evaluations = 24 85
Pair time (%) = 0.776044 (93.833)
Neigh time (%) = 0 (0)
Comm time (%) = 0.0060002 (0.725496)
Outpt time (%) = 0.00200007 (0.241833)
Other time (%) = 0.0430041 (5.19971)
Nlocal: 2047 ave 2047 max 2047 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Nghost: 5765 ave 5765 max 5765 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Neighs: 143220 ave 143220 max 143220 min
Histogram: 1 0 0 0 0 0 0 0 0 0
FullNghs: 286440 ave 286440 max 286440 min
Histogram: 1 0 0 0 0 0 0 0 0 0
Total # of neighbors = 286440
Ave neighs/atom = 139.932
Neighbor list builds = 0
Dangerous builds = 0
#variable Ef equal "c_eatoms" computes the final energy of the cell system after the vacancy
#The final energy is stored to the variable Ef
variable Ef equal "c_eatoms"
variable Ev equal (${Ef}-((${No}-1)/${No})*${Ei})
variable Ev equal (-7245.55429-((${No}-1)/${No})*${Ei})
variable Ev equal (-7245.55429-((2048-1)/${No})*${Ei})
variable Ev equal (-7245.55429-((2048-1)/2048)*${Ei})
variable Ev equal (-7245.55429-((2048-1)/2048)*-7250.3671)
#---------------------------------------------
######################################
# SIMULATION DONE
print "All done"
All done
print "Total number of atoms = ${No}"
Total number of atoms = 2048
print "Initial energy of atoms = ${Ei}"
Initial energy of atoms = -7250.3671
print "Final energy of atoms = ${Ef}"
Final energy of atoms = -7245.55429
print "Vacancy formation energy = ${Ev}"
Vacancy formation energy = 1.272591689
Post-Processing
Visualization
The simulation can be visualized using OVITO (developer: Alexander Stukowski). To do so, first open the Ovito program and select the import data tab. Then go to the directory and file in which you ran the simulation. Locate the first dumpfile, which would be dump.relax.1.0 in this case. You can then double click on the file, or select the file, then click open. The image of the cell should appear on the screen, displayed in four views, Figure 1. This image will be that of the relaxed cell before introducing the vacancy. To view the cell after vacancy has been created, follow the same steps above to select the dump file at the end of the relaxation which followed the atom removal (these files are labeled dump.relax.2.*. Once you open the new file, select one of the four views. Then open the add modification tab and from the drop down menu select Slice. Set the Normal (X) to 1 and set Normal (Y) and Normal (Z) to 0. Return to the add modification tab and select Color coding. When here select c_eng for the Property. Then select Rainbow for the Color gradient and click Adjust range. The vacancy should be visible in the center of the cell, colored by the energy surrounding the vacancy, see Figure 2.
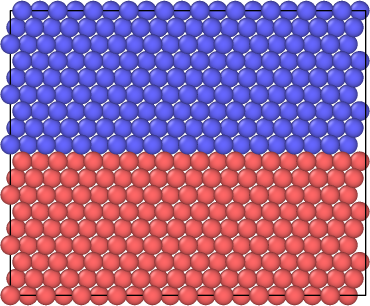

Figure 1: Perspective view of the simulation cell with 2048 total atoms before a vacancy is created.
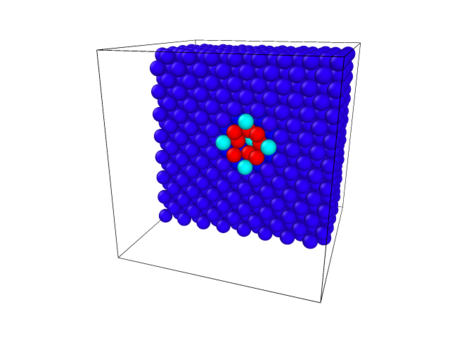
Figure 2: Slice through the center of the simulation cell showing the vacancy. Atoms are colored by the energy per atom. The 8 red atoms are the ones whose bonds were broken with the atom that was removed to create the vacancy, they display the highest energy. The 5 light blue atoms have a lower energy then the red because they were not bonded with the atom that was removed but they shifted as a result of the vacancy due to relaxation
Acknowledgements
We would like to acknowledge the support to this work by the National Science Foundation, HBCUUP-RIA program, Program Manager Dr. Claudia Rankins, Award No. HRD-1137587. Additionally, the technical and logistical support of CAVS and HPC2 of Mississippi State University is gratefully acknowledged.
References
- ↑ S. Plimpton, "Fast Parallel Algorithms for Short-Range Molecular Dynamics," J. Comp. Phys., 117, 1-19 (1995). (http://www.sciencedirect.com/science/article/pii/S002199918571039X)